# VSHunter
**Repository Path**: ShixiangWang/VSHunter
## Basic Information
- **Project Name**: VSHunter
- **Description**: No description available
- **Primary Language**: Unknown
- **License**: MIT
- **Default Branch**: master
- **Homepage**: None
- **GVP Project**: No
## Statistics
- **Stars**: 0
- **Forks**: 0
- **Created**: 2021-08-31
- **Last Updated**: 2024-05-30
## Categories & Tags
**Categories**: Uncategorized
**Tags**: None
## README
# VSHunter: Capture Variation Signatures from Genomic Data
[](https://github.com/ShixiangWang/VSHunter)
[](https://www.tidyverse.org/lifecycle/#retire)
[](https://travis-ci.org/ShixiangWang/VSHunter)
[](https://codecov.io/gh/ShixiangWang/VSHunter) [](https://zenodo.org/badge/latestdoi/153238002)
**This package retired, users should switch to R package [**sigminer**](https://github.com/ShixiangWang/sigminer)**
The goal of VSHunter is to capture variation signature from genomic
data. For now, we decode copy number pattern from **absolute copy number
profile**. This package collects R code from paper *[Copy number
signatures and mutational processes in ovarian
carcinoma](https://www.nature.com/articles/s41588-018-0179-8)* and tidy
them as a open source R package for bioinformatics community.
Before you use this tool, you have to obtain **absolute copy number
profile** for samples via software like ABSOLUTE v2, QDNASeq etc..
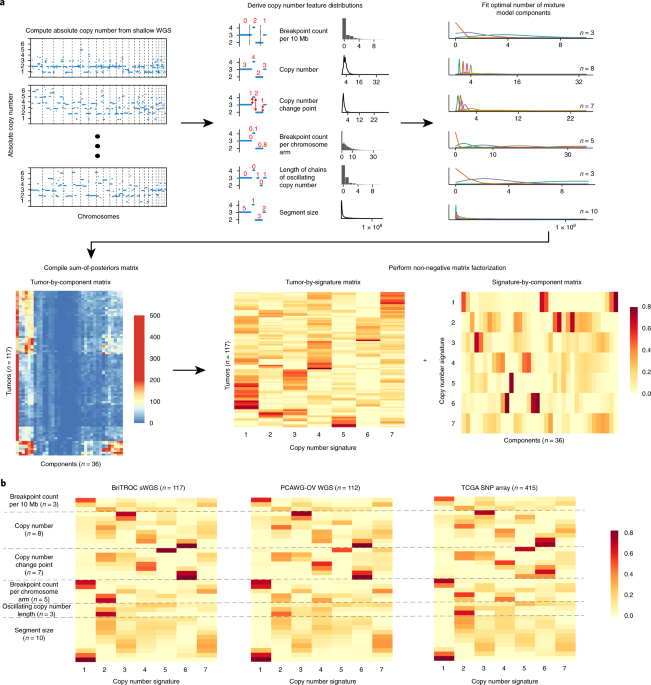
## Procedure
1. summarise copy-number profile using a number of different feature
distributions:
- Sgement size
- Breakpoint number (per ten megabases)
- change-point copy-number
- Breakpoint number (per chromosome arm)
- Length of segments with oscillating copy-number
2. apply mixture modelling to breakdown each feature distribution into
mixtures of Gaussian or mixtures of Poisson distributions using the
flexmix package.
3. generate a sample-by-component matrix representing the sum of
posterior probabilities of each copy-number event being assigned to
each component.
4. use NMF package to factorise the sample-by-component matrix into a
signature-by-sample matrix and component-by
signature-matrix.
 ## Installation
You can install UCSCXenaTools from github with:
``` r
# install.packages("devtools")
devtools::install_github("ShixiangWang/VSHunter", build_vignettes = TRUE)
```
> update features and function will show in vignettes in the future
Load package.
``` r
library(VSHunter)
```
## Citation
- *Macintyre, Geoff, et al. “Copy number signatures and mutational
processes in ovarian carcinoma.” Nature genetics 50.9 (2018): 1262.*
If you wanna thank my work for this package, you can also cite (and
inlucde link of this package -
):
- *Wang, Shixiang, et al. “APOBEC3B and APOBEC mutational signature as
potential predictive markers for immunotherapy response in non-small
cell lung cancer.” Oncogene (2018).*
## Installation
You can install UCSCXenaTools from github with:
``` r
# install.packages("devtools")
devtools::install_github("ShixiangWang/VSHunter", build_vignettes = TRUE)
```
> update features and function will show in vignettes in the future
Load package.
``` r
library(VSHunter)
```
## Citation
- *Macintyre, Geoff, et al. “Copy number signatures and mutational
processes in ovarian carcinoma.” Nature genetics 50.9 (2018): 1262.*
If you wanna thank my work for this package, you can also cite (and
inlucde link of this package -
):
- *Wang, Shixiang, et al. “APOBEC3B and APOBEC mutational signature as
potential predictive markers for immunotherapy response in non-small
cell lung cancer.” Oncogene (2018).* ## Installation
You can install UCSCXenaTools from github with:
``` r
# install.packages("devtools")
devtools::install_github("ShixiangWang/VSHunter", build_vignettes = TRUE)
```
> update features and function will show in vignettes in the future
Load package.
``` r
library(VSHunter)
```
## Citation
- *Macintyre, Geoff, et al. “Copy number signatures and mutational
processes in ovarian carcinoma.” Nature genetics 50.9 (2018): 1262.*
If you wanna thank my work for this package, you can also cite (and
inlucde link of this package -
## Installation
You can install UCSCXenaTools from github with:
``` r
# install.packages("devtools")
devtools::install_github("ShixiangWang/VSHunter", build_vignettes = TRUE)
```
> update features and function will show in vignettes in the future
Load package.
``` r
library(VSHunter)
```
## Citation
- *Macintyre, Geoff, et al. “Copy number signatures and mutational
processes in ovarian carcinoma.” Nature genetics 50.9 (2018): 1262.*
If you wanna thank my work for this package, you can also cite (and
inlucde link of this package -