# Physics-aware-Multiplex-GNN
**Repository Path**: ahlih_admin/Physics-aware-Multiplex-GNN
## Basic Information
- **Project Name**: Physics-aware-Multiplex-GNN
- **Description**: No description available
- **Primary Language**: Unknown
- **License**: Not specified
- **Default Branch**: main
- **Homepage**: None
- **GVP Project**: No
## Statistics
- **Stars**: 0
- **Forks**: 0
- **Created**: 2023-11-22
- **Last Updated**: 2023-11-22
## Categories & Tags
**Categories**: Uncategorized
**Tags**: None
## README
# PAMNet: A Universal Framework for Accurate and Efficient Geometric Deep Learning of Molecular Systems
[](https://paperswithcode.com/sota/drug-discovery-on-qm9?p=a-universal-framework-for-accurate-and)
[](https://paperswithcode.com/sota/protein-ligand-affinity-prediction-on-pdbbind?p=a-universal-framework-for-accurate-and)
Official implementation of **PAMNet** (Physics-aware Multiplex Graph Neural Network) in our paper **[A universal framework for accurate and efficient geometric deep learning of molecular systems](https://www.nature.com/articles/s41598-023-46382-8)** accepted by *Nature Scientific Reports* (doi: 10.1038/s41598-023-46382-8).
PAMNet is an improved version of [MXMNet](https://github.com/zetayue/MXMNet) and outperforms state-of-the-art baselines regarding both accuracy and efficiency in diverse tasks including **small molecule property prediction**, **RNA 3D structure prediction**, and **protein-ligand binding affinity prediction**.
This implementation is also applicable to:
1. Our preprint [Efficient and Accurate Physics-aware Multiplex Graph Neural Networks for 3D Small Molecules and Macromolecule Complexes](https://arxiv.org/abs/2206.02789).
2. Our paper [Physics-aware Graph Neural Network for Accurate RNA 3D Structure Prediction](https://arxiv.org/abs/2210.16392) on [Machine Learning for Structural Biology Workshop](https://www.mlsb.io/) at *NeurIPS 2022*.
If you have any questions, feel free to open an issue or reach out to: szhang4@gradcenter.cuny.edu.
## Overall Architecture
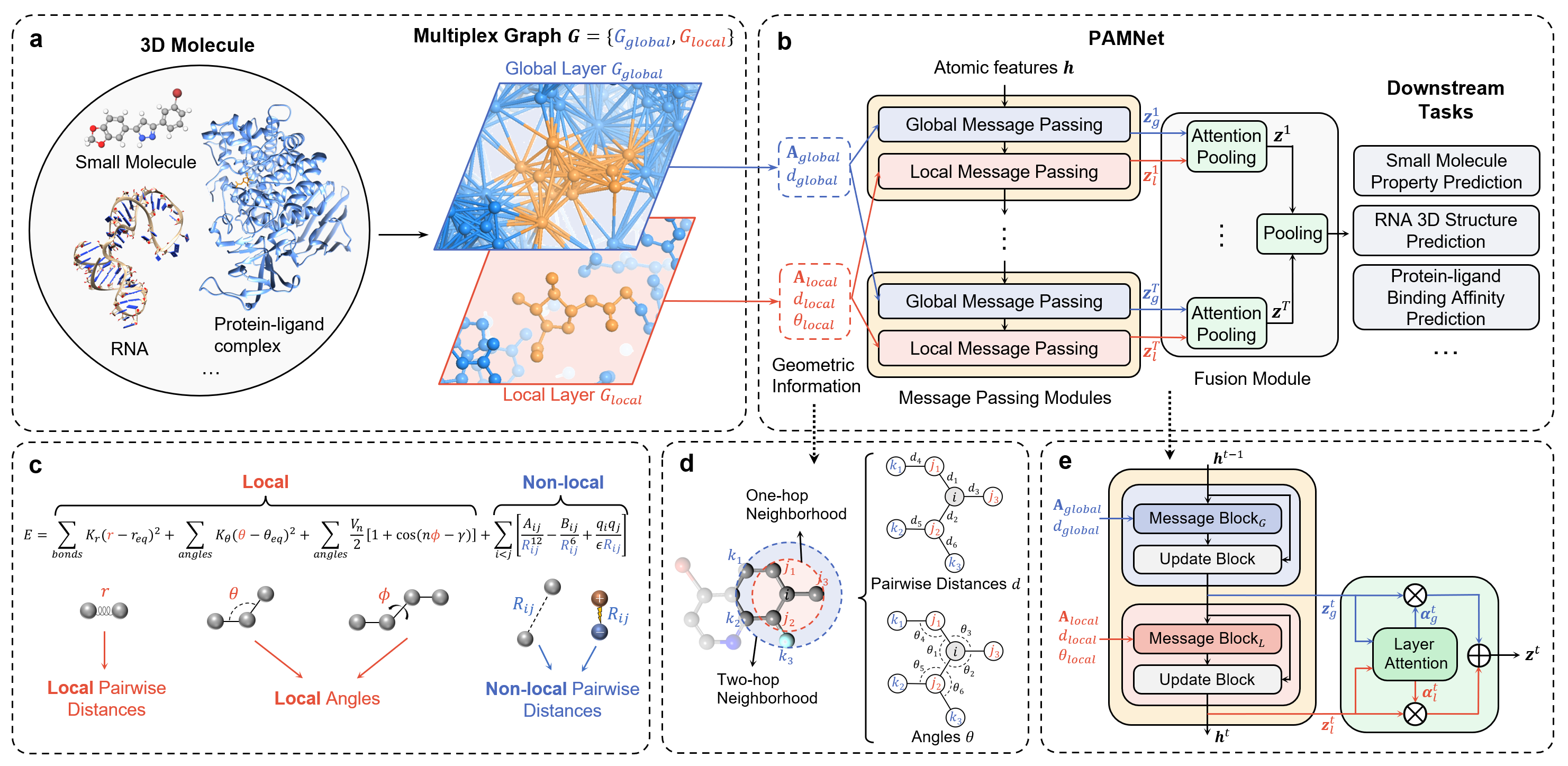
## Setup
### Environment
- Python : 3.7.4
- CUDA : 10.1
**Optional**: Install Open Babel 3.1.1 for binding affinity prediction on PDBbind:
1. Download [source file](https://anaconda.org/conda-forge/openbabel/3.1.1/download/linux-64/openbabel-3.1.1-py37h200e996_1.tar.bz2)
2. `conda install filename`
The other dependencies can be installed with:
```
pip install -r requirements.txt
```
### Datasets
**QM9 for small molecule property prediction:**
The training script (`main_qm9.py`) will automatically download the QM9 dataset and preprocess it.
**PDBbind for protein-ligand binding affinity prediction:**
1. Download `PDBbind_dataset.tar.gz` from [dropbox](https://www.dropbox.com/sh/2uih3c6fq37qfli/AAD-LHXSWMLAuGWzcQLk5WI3a)
2. Unzip the downloaded file under `./data/PDBbind`. There will be two subfolders (`core-set` and `refined-set`) after the unzip
3. Run `python preprocess_pdbbind.py` to preprocess the dataset to construct graphs
**RNA-Puzzles for RNA 3D structure prediction:**
1. Download `classics_train_val.tar` from [Stanford Digital Repository](https://doi.org/10.25740/bn398fc4306)
2. Unzip the downloaded file under `./data/RNA-Puzzles`. There will be one subfolder `classics_train_val` containing `example_train` and `example_val`after the unzip
3. Run `python preprocess_rna_puzzles.py` to preprocess the dataset to construct graphs
## How to Run
### Arguments
```
--gpu GPU number
--seed random seed
--dataset dataset to be used
--epochs number of epochs to train
--lr initial learning rate
--wd weight decay value
--n_layer number of hidden layers
--dim size of input hidden units
--batch_size batch size
--cutoff_l distance cutoff used in the local layer
--cutoff_g distance cutoff used in the global layer
--model model to be used on QM9
--target index of target (0~11) for prediction on QM9
```
### Example command for training and evaluation
**Small molecule property prediction on QM9:**
python -u main_qm9.py --dataset 'QM9' --model 'PAMNet' --target=7 --epochs=900 --batch_size=32 --dim=128 --n_layer=6 --lr=1e-4
**Protein-ligand binding affinity prediction on PDBbind:**
python -u main_pdbbind.py --dataset 'PDBbind' --epochs=170 --batch_size=32 --dim=128 --n_layer=3 --lr=1e-3
**RNA 3D structure prediction on RNA-Puzzles:**
python -u main_rna_puzzles.py --dataset 'RNA-Puzzles' --epochs=15 --batch_size=8 --dim=16 --n_layer=1 --lr=1e-4
## Citation
If you find our model and code helpful in your work, please consider citing us:
```
@article{zhang2023universal,
title={A Universal Framework for Accurate and Efficient Geometric Deep Learning of Molecular Systems},
author={Zhang, Shuo and Liu, Yang and Xie, Lei},
journal={Scientific Reports},
volume={13},
number={1},
pages={19171},
year={2023},
publisher={Nature Publishing Group UK London}
}
@article{zhang2022physics,
title={Physics-aware graph neural network for accurate RNA 3D structure prediction},
author={Zhang, Shuo and Liu, Yang and Xie, Lei},
journal={arXiv preprint arXiv:2210.16392},
year={2022}
}
```
