# single_cell
**Repository Path**: mo_xinwu/single_cell
## Basic Information
- **Project Name**: single_cell
- **Description**: 单细胞相关
- **Primary Language**: R
- **License**: GPL-3.0
- **Default Branch**: master
- **Homepage**: None
- **GVP Project**: No
## Statistics
- **Stars**: 1
- **Forks**: 0
- **Created**: 2023-03-20
- **Last Updated**: 2024-10-10
## Categories & Tags
**Categories**: Uncategorized
**Tags**: None
## README
# README
## 仓库用途
主要用于存放一些单细胞相关的 code
## function
1. 用于 tsne 和 umap 图美化 -> tsne_umap_beautif.R
 2. 用于获取 barcode 的原始信息,并添加 线粒体基因占比、核糖体基因占比、红细胞基因占比信息:Seurat_get_barcode_info.R
3. 添加 plot_gene_dot 函数,用于在 tsne 或 umap 图上展示表达某个基因的细胞 -> tsne_umap_beautif.R
2. 用于获取 barcode 的原始信息,并添加 线粒体基因占比、核糖体基因占比、红细胞基因占比信息:Seurat_get_barcode_info.R
3. 添加 plot_gene_dot 函数,用于在 tsne 或 umap 图上展示表达某个基因的细胞 -> tsne_umap_beautif.R
 4. get_gene_set.R 获取基因集数据
4. get_gene_set.R 获取基因集数据
在进行 GSVA 或 GSEA 分析前,我们需要得到基因集数据,常用的基因集数据有 C1、C2、...、C8、H 9个数据集,可以通过 msigdbr R packages 来获取不同物种的不同数据集。
 - 可用的 Species
| species_name | species_common_name |
| ------------------------------- | ------------------------------------------------------------ |
| Anolis carolinensis | Carolina anole, green anole |
| Bos taurus | bovine, cattle, cow, dairy cow, domestic cattle, domestic cow, ox, oxen |
| Caenorhabditis elegans | NA |
| Canis lupus familiaris | dog, dogs |
| Danio rerio | leopard danio, zebra danio, zebra fish, zebrafish |
| Drosophila melanogaster | fruit fly |
| Equus caballus | domestic horse, equine, horse |
| Felis catus | cat, cats, domestic cat |
| Gallus gallus | bantam, chicken, chickens, Gallus domesticus |
| Homo sapiens | human |
| Macaca mulatta | rhesus macaque, rhesus macaques, Rhesus monkey, rhesus monkeys |
| Monodelphis domestica | gray short-tailed opossum |
| Mus musculus | house mouse, mouse |
| Ornithorhynchus anatinus | duck-billed platypus, duckbill platypus, platypus |
| Pan troglodytes | chimpanzee |
| Rattus norvegicus | brown rat, Norway rat, rat, rats |
| Saccharomyces cerevisiae | baker's yeast, brewer's yeast, S. cerevisiae |
| Schizosaccharomyces pombe 972h- | NA |
| Sus scrofa | pig, pigs, swine, wild boar |
| Xenopus tropicalis | tropical clawed frog, western clawed frog |
5. 双细胞预测: run_scrublet.py
- 可用的 Species
| species_name | species_common_name |
| ------------------------------- | ------------------------------------------------------------ |
| Anolis carolinensis | Carolina anole, green anole |
| Bos taurus | bovine, cattle, cow, dairy cow, domestic cattle, domestic cow, ox, oxen |
| Caenorhabditis elegans | NA |
| Canis lupus familiaris | dog, dogs |
| Danio rerio | leopard danio, zebra danio, zebra fish, zebrafish |
| Drosophila melanogaster | fruit fly |
| Equus caballus | domestic horse, equine, horse |
| Felis catus | cat, cats, domestic cat |
| Gallus gallus | bantam, chicken, chickens, Gallus domesticus |
| Homo sapiens | human |
| Macaca mulatta | rhesus macaque, rhesus macaques, Rhesus monkey, rhesus monkeys |
| Monodelphis domestica | gray short-tailed opossum |
| Mus musculus | house mouse, mouse |
| Ornithorhynchus anatinus | duck-billed platypus, duckbill platypus, platypus |
| Pan troglodytes | chimpanzee |
| Rattus norvegicus | brown rat, Norway rat, rat, rats |
| Saccharomyces cerevisiae | baker's yeast, brewer's yeast, S. cerevisiae |
| Schizosaccharomyces pombe 972h- | NA |
| Sus scrofa | pig, pigs, swine, wild boar |
| Xenopus tropicalis | tropical clawed frog, western clawed frog |
5. 双细胞预测: run_scrublet.py
参数说明:
--gene_cell_matrix_input_file: 输入的 gene x cell矩阵文件行为基因, 列为细胞
--expected_doublet_rate:预估的双细胞率, 默认0.06
--predicted_output_file:结果输出文件, `predicted_doublets` 列为1的即为双细胞,应去除。
|Barcode |doublet_scores |predicted_doublets|
|------------------|--------------------|------------------|
|AAACCCACAAGGCGTA-1|0.04035002430724357 |0 |
|AAACCCACATGGGATG-1|0.031759656652360524|0 |
|AAACCCAGTACTCCGG-1|0.09191277946453216 |0 |
|AAACCCACAGCAGATG-1|0.31719128329297813 |1 |
|AAAGAACAGCTGTACT-1|0.27234342012667123 |1 |
|AAAGTCCAGCTCCACG-1|0.46285714285714286 |1 |
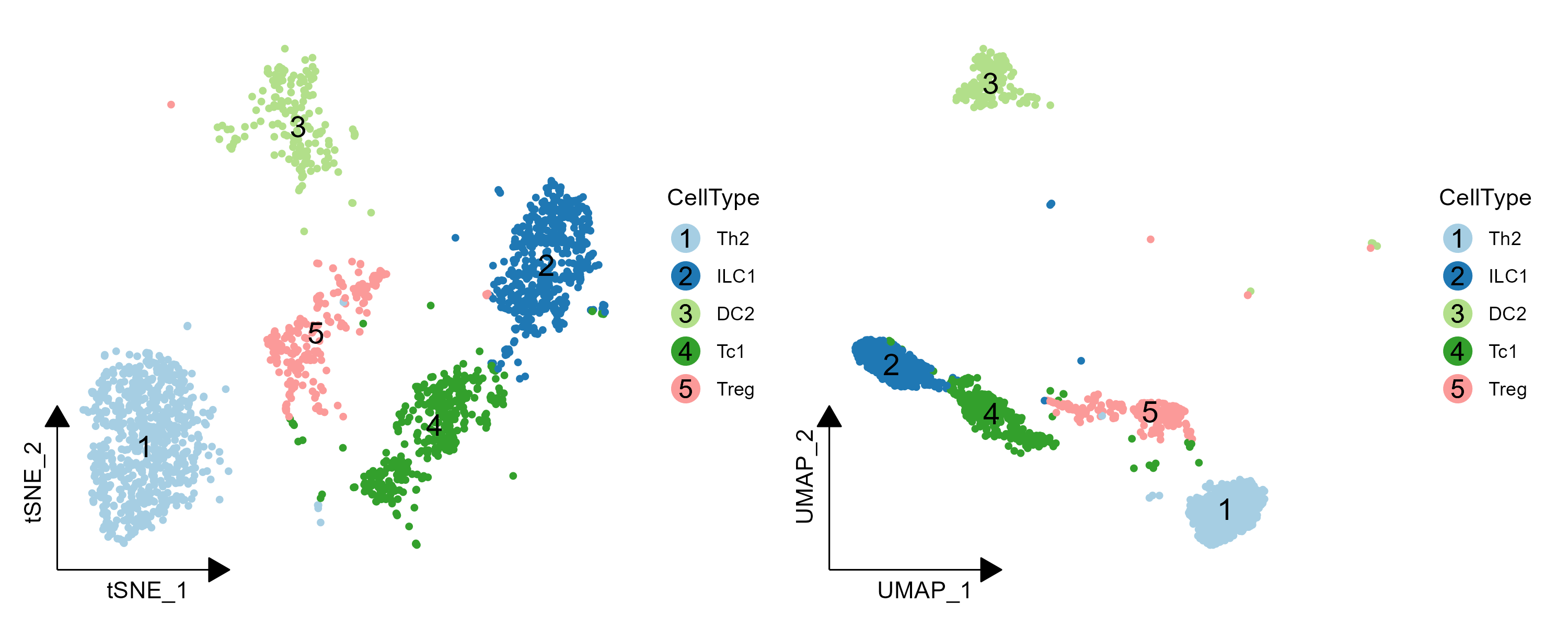
6. legend 为圆圈数字
在 `tsne_umap_beautif.R` 添加 `plot_number_circle` 用于绘制 legend 为圆圈数字, 注意:
- 需要对圆圈进行修大小修改 可以修改 `draw_number_circle`
- 只能用于绘制 一个 cluster 对应 一个 CellType 的情况
- 一定要注意的是 CellType 一定要记得将其转成 factor, 注意顺序, 一定要将 CellType 的 factor 顺序要和 Cluster 的顺序要对对应
```R
data <- data.table::fread("demo_tsne_umap.txt") %>% filter(Cluster %in% c(1,2,3,4,5))
data$CellType <- NA
data$CellType[data$Cluster == "1"] <- "Th2"
data$CellType[data$Cluster == "2"] <- "ILC1"
data$CellType[data$Cluster == "3"] <- "DC2"
data$CellType[data$Cluster == "4"] <- "Tc1"
data$CellType[data$Cluster == "5"] <- "Treg"
data$CellType <- factor(data$CellType,levels = unique(data$CellType)[order(unique(data$Cluster))])
p1 <- plot_number_circle(reduction_data = data,colors_by = "CellType",
labels_by = "Cluster",show_labels = T,
point_size = 1,adjust_axis = T,
label_size = 5)
data[,c(2,3)] <- NULL
p2 <- plot_number_circle(reduction_data = data,colors_by = "CellType",
labels_by = "Cluster",show_labels = T,
point_size = 1,adjust_axis = T,
label_size = 5)
p <- p1 | p2
```
 2. 用于获取 barcode 的原始信息,并添加 线粒体基因占比、核糖体基因占比、红细胞基因占比信息:Seurat_get_barcode_info.R
3. 添加 plot_gene_dot 函数,用于在 tsne 或 umap 图上展示表达某个基因的细胞 -> tsne_umap_beautif.R
2. 用于获取 barcode 的原始信息,并添加 线粒体基因占比、核糖体基因占比、红细胞基因占比信息:Seurat_get_barcode_info.R
3. 添加 plot_gene_dot 函数,用于在 tsne 或 umap 图上展示表达某个基因的细胞 -> tsne_umap_beautif.R
 4. get_gene_set.R 获取基因集数据
4. get_gene_set.R 获取基因集数据  - 可用的 Species
| species_name | species_common_name |
| ------------------------------- | ------------------------------------------------------------ |
| Anolis carolinensis | Carolina anole, green anole |
| Bos taurus | bovine, cattle, cow, dairy cow, domestic cattle, domestic cow, ox, oxen |
| Caenorhabditis elegans | NA |
| Canis lupus familiaris | dog, dogs |
| Danio rerio | leopard danio, zebra danio, zebra fish, zebrafish |
| Drosophila melanogaster | fruit fly |
| Equus caballus | domestic horse, equine, horse |
| Felis catus | cat, cats, domestic cat |
| Gallus gallus | bantam, chicken, chickens, Gallus domesticus |
| Homo sapiens | human |
| Macaca mulatta | rhesus macaque, rhesus macaques, Rhesus monkey, rhesus monkeys |
| Monodelphis domestica | gray short-tailed opossum |
| Mus musculus | house mouse, mouse |
| Ornithorhynchus anatinus | duck-billed platypus, duckbill platypus, platypus |
| Pan troglodytes | chimpanzee |
| Rattus norvegicus | brown rat, Norway rat, rat, rats |
| Saccharomyces cerevisiae | baker's yeast, brewer's yeast, S. cerevisiae |
| Schizosaccharomyces pombe 972h- | NA |
| Sus scrofa | pig, pigs, swine, wild boar |
| Xenopus tropicalis | tropical clawed frog, western clawed frog |
5. 双细胞预测: run_scrublet.py
- 可用的 Species
| species_name | species_common_name |
| ------------------------------- | ------------------------------------------------------------ |
| Anolis carolinensis | Carolina anole, green anole |
| Bos taurus | bovine, cattle, cow, dairy cow, domestic cattle, domestic cow, ox, oxen |
| Caenorhabditis elegans | NA |
| Canis lupus familiaris | dog, dogs |
| Danio rerio | leopard danio, zebra danio, zebra fish, zebrafish |
| Drosophila melanogaster | fruit fly |
| Equus caballus | domestic horse, equine, horse |
| Felis catus | cat, cats, domestic cat |
| Gallus gallus | bantam, chicken, chickens, Gallus domesticus |
| Homo sapiens | human |
| Macaca mulatta | rhesus macaque, rhesus macaques, Rhesus monkey, rhesus monkeys |
| Monodelphis domestica | gray short-tailed opossum |
| Mus musculus | house mouse, mouse |
| Ornithorhynchus anatinus | duck-billed platypus, duckbill platypus, platypus |
| Pan troglodytes | chimpanzee |
| Rattus norvegicus | brown rat, Norway rat, rat, rats |
| Saccharomyces cerevisiae | baker's yeast, brewer's yeast, S. cerevisiae |
| Schizosaccharomyces pombe 972h- | NA |
| Sus scrofa | pig, pigs, swine, wild boar |
| Xenopus tropicalis | tropical clawed frog, western clawed frog |
5. 双细胞预测: run_scrublet.py 